Korean Plant Genetic Coefficient Inference Engine Design
Korean Plant Genetic Coefficient Inference Engine Design
1Core Role Definition (The "Solver")
Forward vs Reverse Process
Forward
Genotype + Environment + Management = Phenotype
Standard prediction structure of crop models
Reverse Engineering
Phenotype + Environment + Management → Genotype
Inverse process of deriving genetic coefficients from observed data
Role of the Inference Engine
The inference engine serves as an Optimizer that minimizes error through tens of thousands of simulations. It iteratively adjusts genetic coefficients until the difference between observed data and simulation results is minimized.
2Specific Construction Steps
Step 1: Simulation Kernel Integration
The 'heart' of the inference engine is the model that simulates actual crop growth.
Crop Model Selection
- Embed the core of validated DSSAT models inside the engine
- Wrap in C# and call via API
Role
Must be able to rapidly compute: "If genetic coefficient A is input, yield B is produced."
Step 2: Search Algorithm Implementation
This is the core engine responsible for 'inference'. The following algorithms are used to find the optimal coefficient combination for Korean cultivars.
Search Algorithm A: Genetic Algorithm (GA)
Finds optimal genetic coefficients by mimicking natural selection.
How It Works
- Generate random sets of genetic coefficients (population)
- Run simulations
- Eliminate high-error individuals; crossover/mutate low-error ones
- Repeat
Suitability
Most commonly used for nonlinear and complex problems like crop models.
Search Algorithm B: Bayesian Inference (MCMC)
Rather than finding a single answer, it infers the probability distribution of possible genetic coefficients.
Advantage
When data is scarce, it can calculate the uncertainty of coefficients alongside, increasing reliability. (GLUE methodology, etc.)
Search Algorithm C: Surrogate Model AI Inference
When physics models (e.g., DSSAT) are slow, a deep learning model (DNN) is pre-trained on the input-output relationships of the physics model.
Training Phase
Run the physics model thousands of times to train a DNN on the input (genetic coefficients) → output (growth results) relationship
Inference Phase
Use the trained DNN to instantly estimate genetic coefficients via backpropagation
Step 3: Objective Function Design
The criterion by which the inference engine judges "how close to the correct answer" it is.
Loss Function
Designed to minimize RMSE (Observed − Predicted).
Target Variables
- Flowering date
- Maturity date
- Yield
- Leaf Area Index (LAI)
3Specialized Strategy for Korean Cultivars
Why is a Specialized Strategy Needed?
To prevent the engine from producing erroneous values (biologically impossible), Korean context must be injected.
Parameter Space Restriction (Bounding Box)
Considering Korea's latitude and climate, upper and lower bounds for coefficients such as photoperiod sensitivity (P1) and grain filling (P5) are preset to narrow the search space.
Cultivar Clustering
Cultivars are grouped by maturity (early, medium, late), and different initial estimates (Priors) are set for each group to improve inference speed and accuracy.
4System Architecture
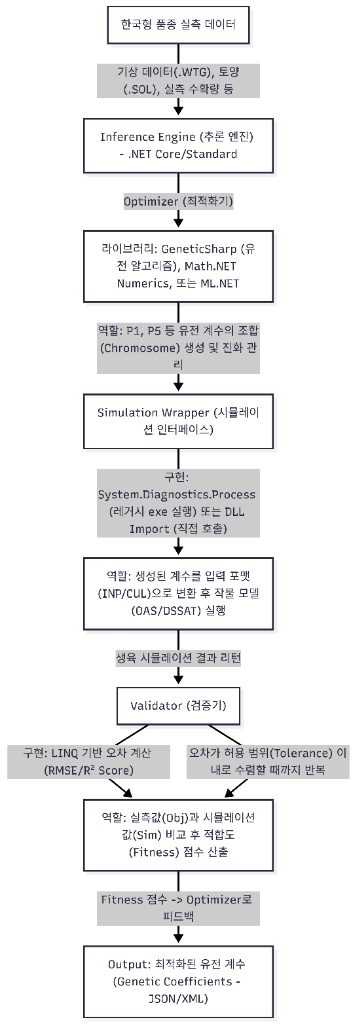
Architecture Components
Observed Cultivar Data
Input data including weather data (WTG), soil (SOL), and observed yield
Inference Engine
.NET Core/Standard based inference engine and Optimizer
Output
Optimized Genetic Coefficients (JSON/XML)
Iterative Process
- Optimizer: Coefficient search using libraries (GeneticSharp, Math.NET Numerics, ML.NET)
- Simulation Wrapper: Call simulation engine (OAS/DSSAT) from .NET
- Validator: Compare observed vs predicted values to compute fitness
- Feedback Loop: Feed back to Optimizer until fitness reaches target
5Summary
Key to Building the Inference Engine
To concretely build this pipeline's inference engine, a crop growth model (Simulator) must be wrapped with an optimization algorithm (Optimizer), creating an iterative loop program that automatically adjusts genetic coefficients until the error between observed data and simulation results is minimized.